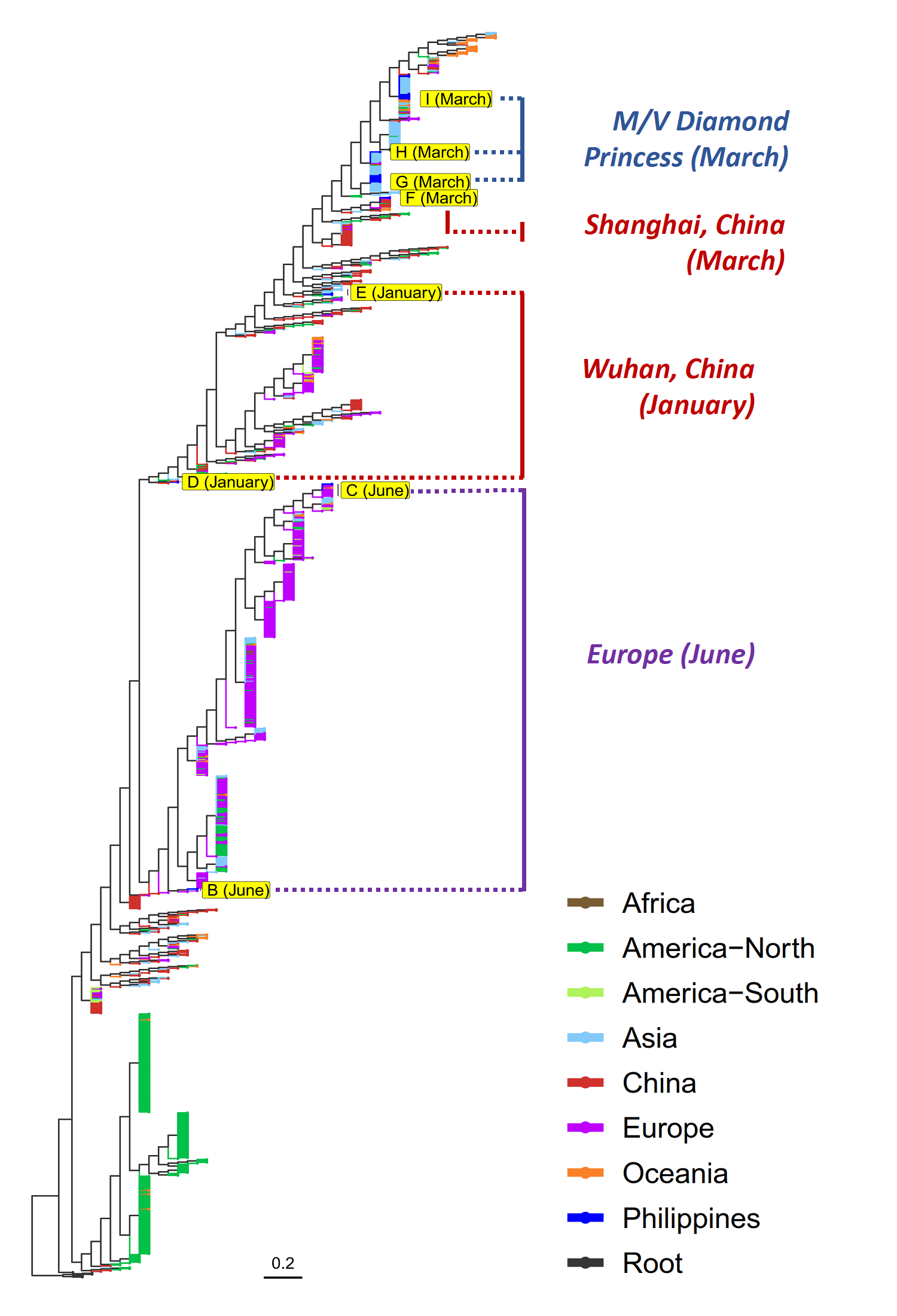
[Findings based on analysis of the whole genome sequences of SARS-CoV-2 submitted to the GISAID (Global Initiative on Sharing All Influenza Data) database by RITM (17) and the PGC (6)]
In a paper posted on August 25, 2020 in the medRxiv1 preprint website, the Philippine Genome Center posits at three possible main sources of COVID-19 infection into the country. PGC’s phylogenomic analysis of 1,335 SARS-CoV-2 sequences in GISAID, including the 23 Philippine submissions, show that the local virus isolates cluster into eight (8) clades that could be further classified into three groups based on possible sources or entry routes of infection: (i) foreign visitors from China with early samples collected in January being closely related to the original isolates from Wuhan, the epicenter of the pandemic at that time; (ii) repatriated seafarers from the M/V Diamond Princess COVID-19 outbreak in Yokohama, Japan with samples collected in March being primarily community transmissions linked to the M/V Diamond Princess cruise ship cluster of cases; and (iii) European sources with samples from June primarily clustering with European isolates, possibly from repatriated tourists and OFWs returning in May from Europe and the Middle East.
Majority of the Philippine submissions (18 of 23) were collected in the month of March, wherein except for one sample which clustered with isolates from Shanghai, China, all others were observed to group into clades linked to the outbreak in the cruise ship, M/V Diamond Princess, moored in Yokohama, Japan in early February 2020. Later that month, passengers and crew members of this cruise ship representing various nationalities including Filipinos, Indians, and Australians were repatriated to their home countries.
Repatriation of a total of 440 Filipino crew members and five tourists from the M/V Diamond Princess on February 26 facilitated the return to the country of the majority of the ship’s crew members who were then quarantined for 14 days at New Clark City in Capas, Tarlac.2 Within 10 days after the end of the quarantine period, we see community transmissions in patients admitted to the Philippine General Hospital whose swab samples were taken between March 22 to 26. All six patients have had no travel history outside the country during this month, with some having no known interaction with a confirmed positive case – suggesting the possibility of community-acquired transmission from undetected cases of infection among the repatriated seafarers.
These findings highlight the need to continually review and improve the quarantine, testing and tracing measures being employed locally to adapt to the current pandemic situation. Nevertheless, the strict imposition of the Enhanced Community Quarantine (ECQ) in the entire island of Luzon, where majority of the confirmed cases originated, from March 17 to May 31, 2020 might have stymied the further spread of the virus from the M/V Diamond Princess cluster of cases, as no such related cases were observed in the months of June and July (cases of which are mostly linked to European clusters). We note, though, that these observations come from only a few local viral isolates with available sequence data and has a limited geographic reach as these were mostly collected in Metro Manila. In addition, while no associations can be made between phylogenomic clustering and disease trajectory, old age still appeared to be a major risk factor in disease severity although co-morbidities also substantially affected clinical outcomes.
The Philippine Genome Center will continue to study through whole genome sequencing and bioinformatics analysis the circulating strains of the SARS-CoV-2 virus in the country to better inform the DOH and the IATF, infectious disease specialists and other stakeholders.
1Tablizo FA et. al. 2020. Analysis of SARS-CoV-2 Genome Sequences from the Philippines: Genetic Surveillance and Transmission Dynamics. medRxiv
doi: doi.org/10.1101/2020.08.22.20180034
This article is a preprint and has not been certified by peer review.
2DOH Press Release/February 26, 2020